High-Level Research Paper Series: Artificial Intelligence in Plant Breeding Uncovers Novel Mechanisms for Crop Breeding
The GSCAAS Pakistan student MUHAMMAD AMJAD FAROOQ (Batch 2023), under the supervision of Professor Li Huihui at the Institute of Crop Sciences of the Chinese Academy of Agricultural Sciences (ICS,CAAS), published an article entitled "Artificial Intelligence in Plant Breeding" as the first author in Trends in Genetics (JCR-Q1, IF = 16.3). This study is a comprehensive review exploring the transformative role of artificial intelligence (AI) across all major facets of modern plant breeding, from automated data collection and high-throughput image-based phenotyping to genomic selection models, gene bank diversity screening, multi-omics data integration, and AI-assisted precision gene editing, providing a roadmap for how AI-enabled breeding can accelerate genetic gain, bridge the genotype-to-phenotype gap, and develop crop cultivars tailored to future environments.
This study was supported by grants from the Sustainable Development International Cooperation Program from the Bill and Melinda Gates Foundation (2022YFAG1002), the National Natural Science Foundation of China (32261143757), Nanfan special project, CAAS (YBXM2407), the Key R&D Programs of Hainan Province (ZDYF2024XDNY210), the Bill and Melinda Gates Foundation (INV-030574) on mining useful alleles for climate change adaptation from CGIAR gene banks, and the Innovation Program of Chinese Academy of Agricultural Sciences (CAAS-CSIAF-202303).
Original link of the article:
https://doi.org/10.1016/j.tig.2024.07.001
Research Introduction
The longstanding assumption that improving crop yields inevitably weakens disease resistance has shaped both evolutionary biology and plant breeding for decades. However, the genetic mechanisms underlying this trade-off remain poorly understood, particularly in complex polyploid crops such as bread wheat. To address this directly, this study constructed an orthology-based pan-genome from 37 high-quality wheat genome assemblies, spanning the entire evolutionary journey of wheat from wild diploid ancestors (including Triticum turgidum subsp. dicoccoides and Aegilops tauschii) through tetraploid forms, to modern hexaploid cultivars.
By tracking the presence and absence of yield- and resistance-related genes (NLR genes) across all 37 genomes, the study uncovered that both gene sets are remarkably well conserved throughout wheat's evolutionary history. Yield-related genes predominantly reside in the stable "core" pan-genome shared by nearly all lines, while resistance genes are more frequently found in the flexible "variable" regions that differ between individual accessions. Despite these contrasting distribution patterns, a strong positive relationship between yield gene count and resistance gene count emerged specifically after polyploidization. This relationship was weak or even negative in wild diploid ancestors.
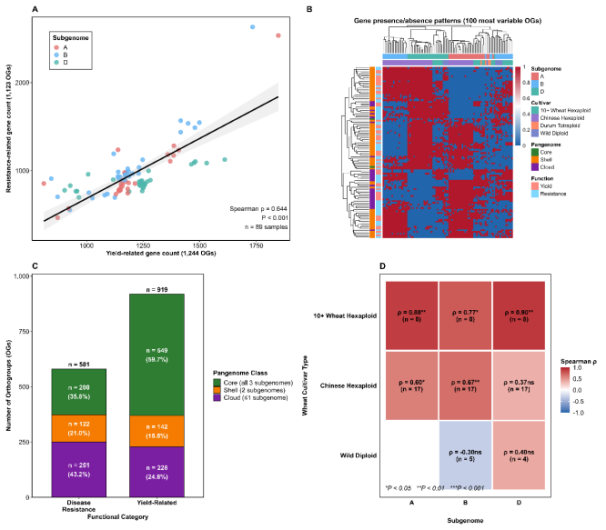
Figure: Pan-genomic analysis of yield- and resistance-related gene relationships across wheat genomes. Panel A: Correlation between yield-related gene count and resistance-related gene count across 89 wheat samples. Panels B: Heatmap of gene presence/absence patterns across the 100 most variable orthogroups. Panel C: Pan-genome class distribution of disease resistance and yield-related orthogroups. Panel D: Heatmap of Spearman correlation coefficients between yield- and resistance-related gene counts across wheat cultivar types.
Sub-genome analysis further identified the B sub-genome as a key contributor to maintaining this balance. Importantly, the findings show that the domestication process did not cause a substantial loss of resistance genes or a reduction in yield, contradicting the long-held assumption that early farmers sacrificed disease-defense mechanisms in pursuit of improved yields. Taken together, these findings show that high yield and strong disease resistance are not fundamentally opposed at the genomic level, and that polyploidy and pan-genome structural diversity enable crops to optimize both traits simultaneously.
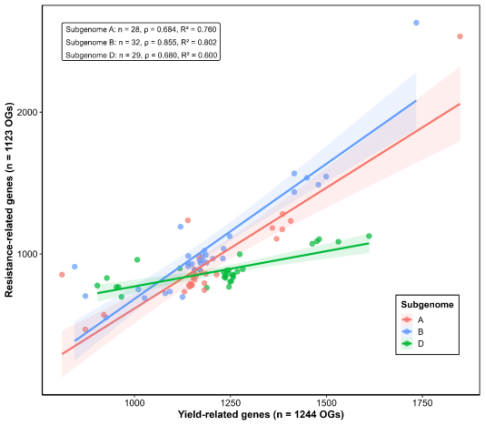
Figure: Sub-genome-specific linear regressions between resistance-related gene counts (1,123 orthogroups) and yield-related gene counts (1,244 orthogroups).
About Author

MUHAMMAD AMJAD FAROOQ is currently in his 3rd year of the master's program at the Institute of Crop Sciences under the supervision of Professor Li Huihui at the Institute of Crop Sciences of the Chinese Academy of Agricultural Sciences (CAAS). His major is Bioinformatics (Research Direction: Big Data Intelligent Breeding). His study in China is financially supported by the GSCAAS Scholarship (Graduate School of the Chinese Academy of Agricultural Sciences). During his studies, he published a research article as first author in Trends in Genetics (JCR-Q1, IF = 16.3), one of the leading international journals in genetics and genomics. In addition, he received Outstanding Volunteer Award from Pakistan Prime Minister's Initiative on Capacity Building for 1,000 Agricultural Graduates and awarded by the National Nanfan Research Institute of CAAS, Yazhou, Sanya, China. Currently, he is also a Project Intern of Gates Fellowship Program, Assisting National Agricultural Research Systems (NARS) experts from Africa at the National Nanfan Research Institute of CAAS, Yazhou, Sanya, China. Supported by the Bill & Melinda Gates Foundation to facilitate international agricultural research and capacity building.
MUHAMMAD AMJAD FAROOQ’s Publications list:
https://doi.org/10.1016/j.tig.2024.07.001
https://doi.org/10.1007/s10722-024-02029-9
About Supervisor

Dr. LI Huihui, Research Professor and Doctoral Supervisor. She serves as Deputy Director of the Center of Crop Genes and Molecular Design (ICS,CAAS), and concurrently holds a professorship at the School of Statistics, Beijing Normal University. She is also Secretary-General of the Smart Agriculture Committee of the Crop Science Society of China, Standing Member of the Agricultural and Forestry Informatics Professional Committee of the Chinese Society of Bioinformatics (in preparation), and a Council Member of the Chinese Society for Tropical Crops. In academic publishing, she serves as Section Editor-in-Chief of Frontiers in Plant Science, and sits on the editorial boards of aBIOTECH, Theoretical and Applied Genetics, Transactions of the Chinese Society of Agricultural Engineering, Frontiers in Genetics, and Acta Agronomica Sinica. A recipient of the National Talent Program and the Leading Talent of Agricultural Science Elite Award from the Chinese Academy of Agricultural Sciences (CAAS), Dr. Li is the Chief Scientist of CAAS’s "Big Data-Driven Intelligent Design Breeding" Innovation Team, and has been honored with the Young Science and Technology Award from the Chinese Association of Agricultural Science Societies.
Her research focuses on the development and application of intelligent algorithm models for crop breeding. As first or corresponding author, she has published 60 papers in top-tier journals including Molecular Plant, Nature Plants, Plant Biotechnology Journal, Trends in Plant Science, and Trends in Genetics. Her work was selected as one of Hainan Province’s Top 10 Outstanding Agricultural Scientific and Technological Achievements in both 2023 and 2024, and she received the Award for Excellent Solutions in Biological Breeding from the China Association for Science and Technology. Moreover, she is the author of the monograph Gene Mapping and Breeding Design, holds 12 authorized patents, and has led or participated in numerous national-level projects, including grants from the National Natural Science Foundation of China and the National Key R&D Program of China.
